An article published 19 January 2016 in Nature Communications, “Iron Age and Anglo-Saxon genomes from East England reveal British migration history,” by Stephan Schiffels et al1, estimates that “the modern-day East English population derives on average 38% of its ancestry from Anglo-Saxon migrants.”
The abstract:
British population history has been shaped by a series of immigrations, including the early Anglo-Saxon migrations after 400 CE. It remains an open question how these events affected the genetic composition of the current British population. Here, we present whole-genome sequences from 10 individuals excavated close to Cambridge in the East of England, ranging from the late Iron Age to the middle Anglo-Saxon period. By analysing shared rare variants with hundreds of modern samples from Britain and Europe, we estimate that on average the contemporary East English population derives 38% of its ancestry from Anglo-Saxon migrations. We gain further insight with a new method, rarecoal, which infers population history and identifies fine-scale genetic ancestry from rare variants. Using rarecoal we find that the Anglo-Saxon samples are closely related to modern Dutch and Danish populations, while the Iron Age samples share ancestors with multiple Northern European populations including Britain.
Last year an equally fascinating study of the genetic structure of the British population (actually UK; while Northern Ireland was included, the Republic of Ireland was not) in Nature last year showed that populations were surprisingly geographically distinct, and they estimate that the Anglo-Saxons contributed a good chunk, though less than half, of the tested populations’ genetic material in the southeast. But as this most recent article points out, inferring these proportions from contemporary data means relying on relatively weak evidence. By contrast, working directly with whole-genome sequences from 10 ancient samples taken from individuals excavated from archaeological contexts ranging from Iron Age to early and middle Anglo-Saxon periods, lets us “directly observe and quantify the genetic impact of the Anglo-Saxon migrations in England.”
Their samples show “mixing of migrants and natives in the early Anglo-Saxon period,” and “the Anglo-Saxon migrants studied here have close ancestry to modern-day Dutch and Danish populations.”
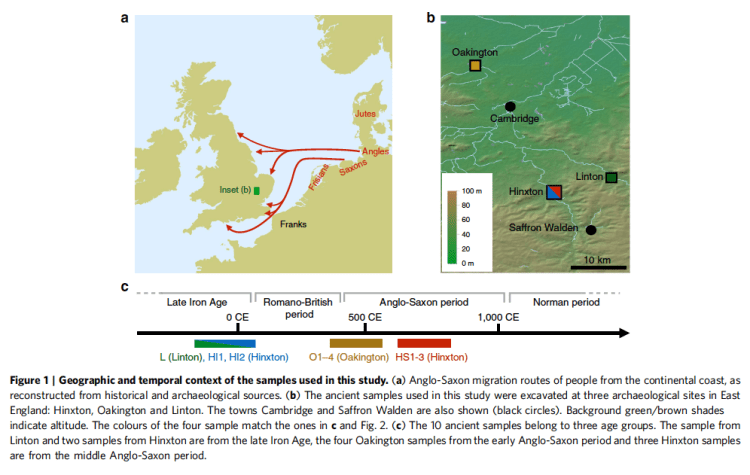
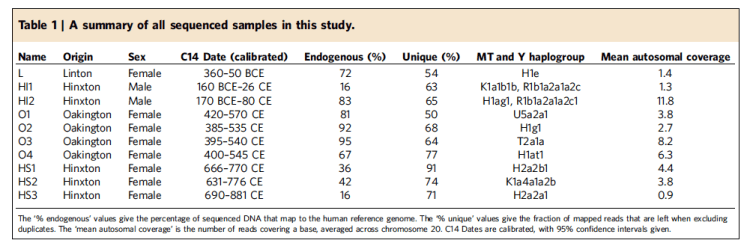
The 10 samples were from three different sites: Linton, Hinxton, and Oakington, all near Cambridge. They were carbon-dated and assigned to three general periods (Iron Age, Early A-S (5t-6th C), middle A-S (7th-9th).
But the specific breakdowns of carbon dating is, I think, worth a look:
And look at how, by identifying “rare variants with allele frequency up to 1% in a reference panel of 433 European individuals from modernFinland, Spain, Italy, Netherlands and Denmark, for which genome-wide sequence data are available” the study “determined for each ancient sample the number of rare variants shared with each reference population.” In other words, how the 10 samples fit into the European genetic tree:
Here are the study’s conclusions (my emphasis):
Our analysis of early and middle Anglo-Saxon samples from East England adds significantly to our picture of the Anglo-Saxon period in Britain. In the cemetery at Oakington we see evidence even in the early Anglo-Saxon period for a genetically mixed but culturally Anglo-Saxon community, in contrast to claims for strong segregation between newcomers and indigenous peoples. The genomes of two sequenced individuals (O1 and O2) are consistent with them being of recent immigrant origin, from a source population close to modern Dutch, one was genetically similar to native Iron Age samples (O4), and the fourth was consistent with being an admixed individual (O3), indicating interbreeding. Despite this, their graves were conspicuously similar, with all four individuals buried in flexed position, and with similar grave furnishing. Interestingly the wealthiest grave, with a large cruciform brooch, belonged to the individual of native British ancestry (O4), and the individual without grave goods was one of the two genetically ‘foreign’ ones (O2), an observation consistent with isotope analysis at West Heslerton which suggests that new immigrants were frequently poorer.
Up to this point we have interpreted the genetic structure of the Anglo-Saxon samples in terms of recent immigrant versus indigenous populations. However, in the absence of a time series through the Romano-British period from the Iron Age to the Anglo-Saxon period, we should also consider the possibility that some of the genetic heterogeneity seen in the Oakington samples arose earlier due to immigration in Romano-British times. We recall that sample O4 lies genetically almost centrally in the Iron Age samples, and O1 and O2 are very close to the later Middle Saxon samples from Hinxton and modern Dutch. For Roman immigration patterns to generate this diverse structure in the fifth to sixth century Oakington samples, one would have to assume strong social segregation with little interbreeding over multiple generations. This seems unlikely given that immigration into Roman-Britain was geographically diverse and consisted of an administrative elite28 and the military, who would have interbred and recruited locally, particularly in the last decades of the third and fourth centuries. Furthermore, there is no significant Roman settlement at Oakington and no evidence for significant Roman Heritage. Given the mixing apparent 500 CE, and that the modern population is not more than 40% of Anglo-Saxon ancestry, it is perhaps surprising that the middle Anglo-Saxon individuals from the more dispersed field cemetery in Hinxton look more genetically consistent with unmixed immigrant ancestry. One possibility is that this reflects continued immigration until at least the Middle Saxon period. The unmixed Hinxton group, versus the mixing of the Oakington population, shows that early medieval migration took a variety of forms and that these migrants integrated with the incumbent population in different ways. Full-genome sequences, and new methods such as rarecoal, now allow us to use slight distinctions in genetic ancestry to study such recent events. Further ancient genomes, and methodological improvements to incorporate explicit migration and mixing, will enable us to resolve them in more detail.
What does this mean in terms of numbers of immigrants? 38% is a lot. Think of it as each south-east English person of, say, the ninth-century having three out of eight great-grandparents of Anglo-Saxon origin.
So did those ancestors come over as large and poor immigrant families with many children? (This mirrors what we’ve seen in last hundred years: the newly-arrived don’t bring much with them.) Were there endless boatloads of fighting men who came for purposes of relentless reproduction? (I consider this less likely.) I don’t have estimates for population numbers in the three periods, but it’s clear that whichever way we slice this we’re talking a significant number of immigrants. If there were only a million Romano-British to start with, and assuming static population numbers (a silly assumption, but we have to start somewhere), we’re still talking nearly 400,000 incomers. Those are big numbers.
Did they come in a steady stream over the centuries or was there the occasional giant wave? Either way, I’m curious about what prompted such mass movements of people.
The most obvious precipitating factor is weather change, either sudden (leading to occasional but massive waves of immigrants) or gradual (steady, long-term influx). Weather change, long- or short-term, leads to disease, and triggers shortages (of land, of food, of water) which lead to conflict over resources. Which of course leads to more shortages, more disease, and more conflict.
So my question is, should we be looking at catastrophic events (meteoroids, volcanoes) or more subtle, cyclical changes (such as El Niño or other natural oscillations)? Right now, if I had to guess, I’d say both, with—of course—interplay between them.
The more we look at ancient genomes, the more fine-grained our understanding of immigration patterns will become. I’d like to venture the hope that ancient remains are also, where possible, subjected to shotgun proteomics so we can begin to get a picture of the diseases to which our ancestors had immune responses.
I’ve been thinking a lot lately about bubonic (and pneumonic, and tonsillar) plague. Also smallpox and malaria. Even measles. I wouldn’t be at all surprised to find evidence of their hand on our ancestry. (Look at what happened when Europeans first arrived in the Americas.) But we can’t know until we look. I hope someone is looking.
ETA: Apparently some of the SNPs (single-nucleotide polymorphisms) associated with smallpox and plague resistance should be available from the genome. So has anyone done this on these samples? Anyone know?
1 Schiffels, S. et al. Iron Age and Anglo-Saxon genomes from East England reveal British migration history. Nat. Commun. 7:10408 doi: 10.1038/ncomms10408 (2016). All quotes and images used under Creative Commons licence.